Joshua S. Weinstock, Jayakrishnan Gopakumar, Bala Bharathi Burugula, Md Mesbah Uddin, Nikolaus Jahn, Julia A. Belk, Hind Bouzid, Bence Daniel, Zhuang Miao, Nghi Ly, Taralynn M. Mack, Sofia E. Luna, Katherine P. Prothro, Shaneice R. Mitchell, Cecelia A. Laurie, Jai G. Broome, Kent D. Taylor, Xiuqing Guo, Moritz F. Sinner, Aenne S. von Falkenhausen, Stefan Kääb, Alan R. Shuldiner, Jeffrey R. O’Connell, Joshua P. Lewis, Eric Boerwinkle, Kathleen C. Barnes, Nathalie Chami, Eimear E. Kenny, Ruth J. F. Loos, Myriam Fornage, Lifang Hou, Donald M. Lloyd-Jones, Susan Redline, Brian E. Cade, Bruce M. Psaty, Joshua C. Bis, Jennifer A. Brody, Edwin K. Silverman, Jeong H. Yun, Dandi Qiao, Nicholette D. Palmer, Barry I. Freedman, Donald W. Bowden, Michael H. Cho, Dawn L. DeMeo, Ramachandran S. Vasan, Lisa R. Yanek, Lewis C. Becker, Sharon L. R. Kardia, Patricia A. Peyser, Jiang He, Michiel Rienstra, Pim Van der Harst, Robert Kaplan, Susan R. Heckbert, Nicholas L. Smith, Kerri L. Wiggins, Donna K. Arnett, Marguerite R. Irvin, Hemant Tiwari, Michael J. Cutler, Stacey Knight, J. Brent Muhlestein, Adolfo Correa, Laura M. Raffield, Yan Gao, Mariza de Andrade, Jerome I. Rotter, Stephen S. Rich, Russell P. Tracy, Barbara A. Konkle, Jill M. Johnsen, Marsha M. Wheeler, J. Gustav Smith, Olle Melander, Peter M. Nilsson, Brian S. Custer, Ravindranath Duggirala, Joanne E. Curran, John Blangero, Stephen McGarvey, L. Keoki Williams, Shujie Xiao, Mao Yang, C. Charles Gu, Yii-Der Ida Chen, Wen-Jane Lee, Gregory M. Marcus, John P. Kane, Clive R. Pullinger, M. Benjamin Shoemaker, Dawood Darbar, Dan M. Roden, Christine Albert, Charles Kooperberg, Ying Zhou, JoAnn E. Manson, Pinkal Desai, Andrew D. Johnson, Rasika A. Mathias, NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium, Thomas W. Blackwell, Goncalo R. Abecasis, Albert V. Smith, Hyun M. Kang, Ansuman T. Satpathy, Pradeep Natarajan, Jacob O. Kitzman, Eric A. Whitsel, Alexander P. Reiner, Alexander G. Bick & Siddhartha Jaiswal
Nature, 12 April 2023
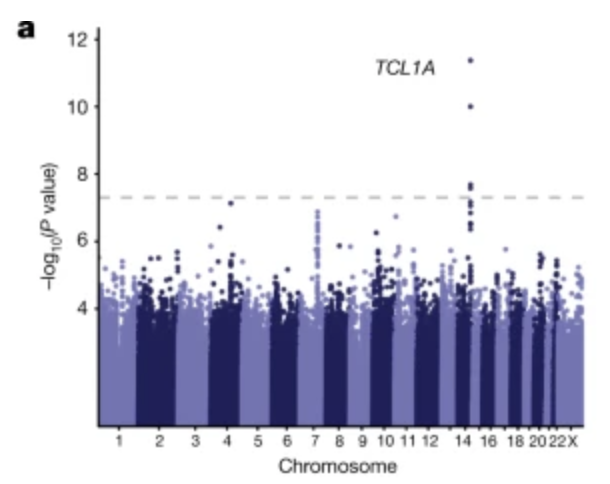
Mutations in a diverse set of driver genes increase the fitness of haematopoietic stem cells (HSCs), leading to clonal haematopoiesis1. These lesions are precursors for blood cancers2-6, but the basis of their fitness advantage remains largely unknown, partly owing to a paucity of large cohorts in which the clonal expansion rate has been assessed by longitudinal sampling. Here, to circumvent this limitation, we developed a method to infer the expansion rate from data from a single time point. We applied this method to 5,071 people with clonal haematopoiesis. A genome-wide association study revealed that a common inherited polymorphism in the TCL1A promoter was associated with a slower expansion rate in clonal haematopoiesis overall, but the effect varied by driver gene. Those carrying this protective allele exhibited markedly reduced growth rates or prevalence of clones with driver mutations in TET2, ASXL1, SF3B1 and SRSF2, but this effect was not seen in clones with driver mutations in DNMT3A. TCL1A was not expressed in normal or DNMT3A-mutated HSCs, but the introduction of mutations in TET2 or ASXL1 led to the expression of TCL1A protein and the expansion of HSCs in vitro. The protective allele restricted TCL1A expression and expansion of mutant HSCs, as did experimental knockdown of TCL1A expression. Forced expression of TCL1A promoted the expansion of human HSCs in vitro and mouse HSCs in vivo. Our results indicate that the fitness advantage of several commonly mutated driver genes in clonal haematopoiesis may be mediated by TCL1A activation.