Research
We use numerous immunological model systems to do so, but we place a particular emphasis on human immunology and bedside-to-bench research. Namely, we study immunological mechanisms of disease directly in samples isolated from patients, and thus, insights gained from this research are likely to be directly applicable to the design of improved immunotherapies for patients.
Projects
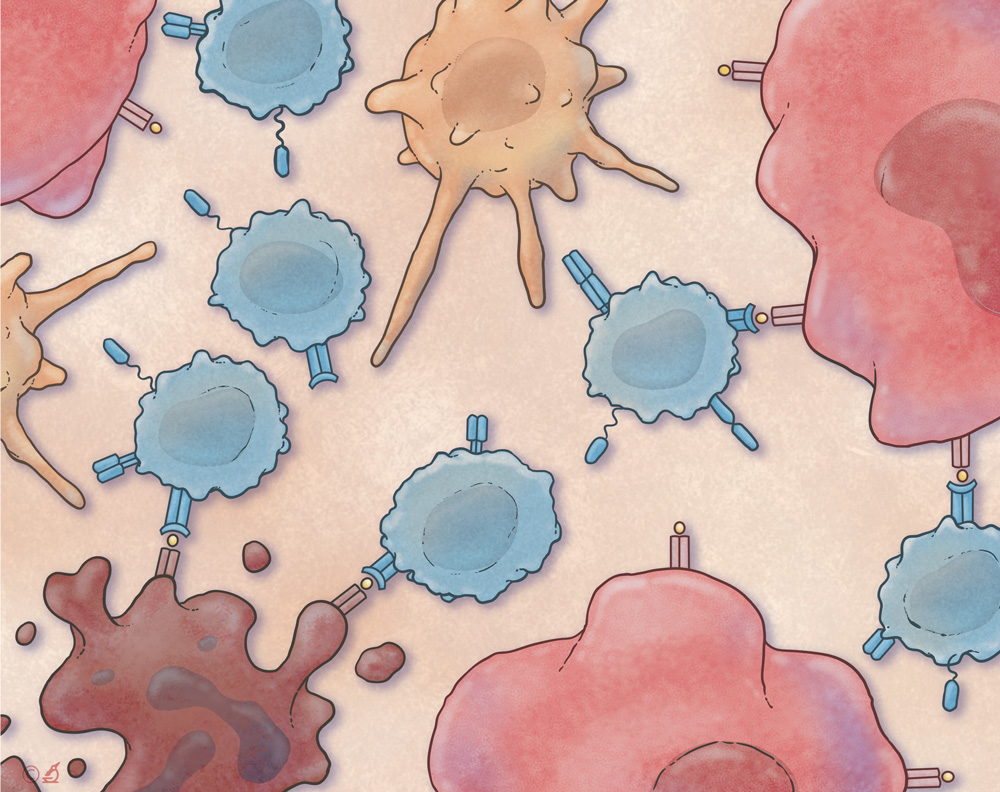
Single-Cell Genomics in Cancer Immunology
Cellular heterogeneity underlies all biological processes, for example, variability in immune responses or cancer relapse rates following immunotherapy. We develop and employ novel single-cell techniques to understand several key questions:
1) What are the cell types that underlie the immune responses to cancer and cancer immunotherapy?
2) What are the genetic pathways that regulate these cellular pathways?
3) How do single cells persist or change over time to promote effective anti-tumor immunity?
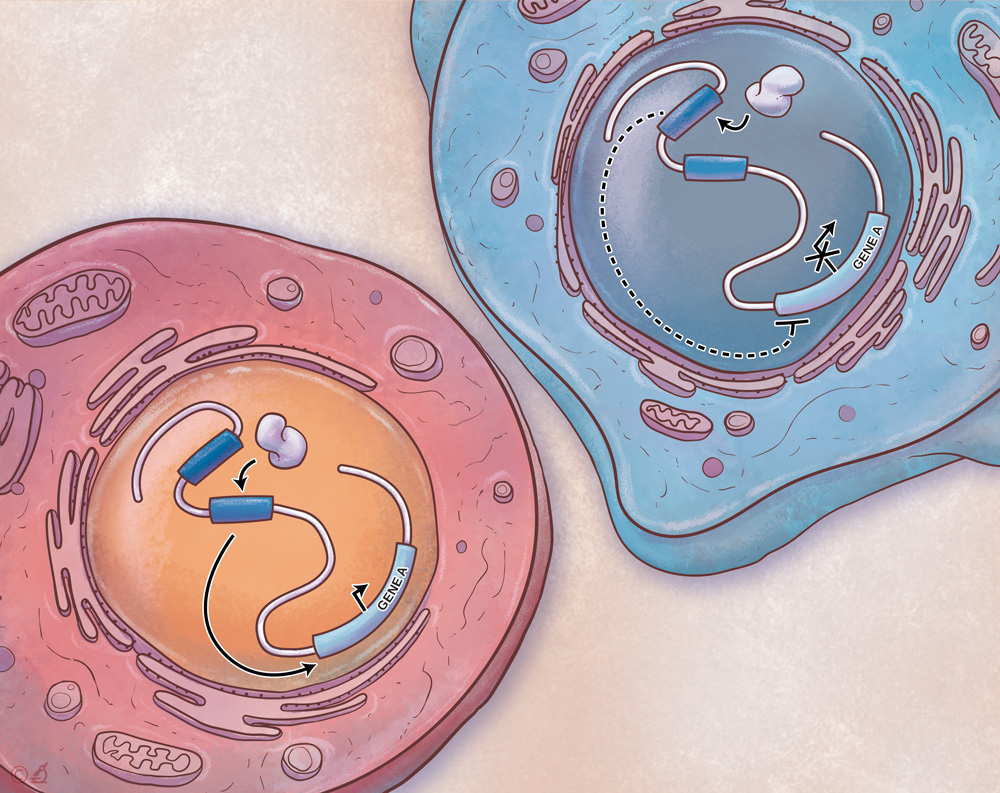
High-throughput Perturbation Technologies for Immune Cells
To functionally interrogate the molecular programs driving key immune cell types and activation states in disease, we utilize a toolbox of high-throughput perturbation technologies using CRISPR/Cas9 genome editing. These technologies enable us to ask several critical questions:
1) What are the key molecular programs (i.e. genes, transcription factors, and enhancer elements) that control key cell type behaviors, such as T cell phenotypes within the tumor microenvironment?
2) Which programs are shared between immune cell types, cancer types, and/or checkpoint blockade agents, and which programs are signal-specific?
3) Can these modules be altered to induce cells states associated with productive immunity, or to reverse dysfunctional cell states associated with therapy resistance?
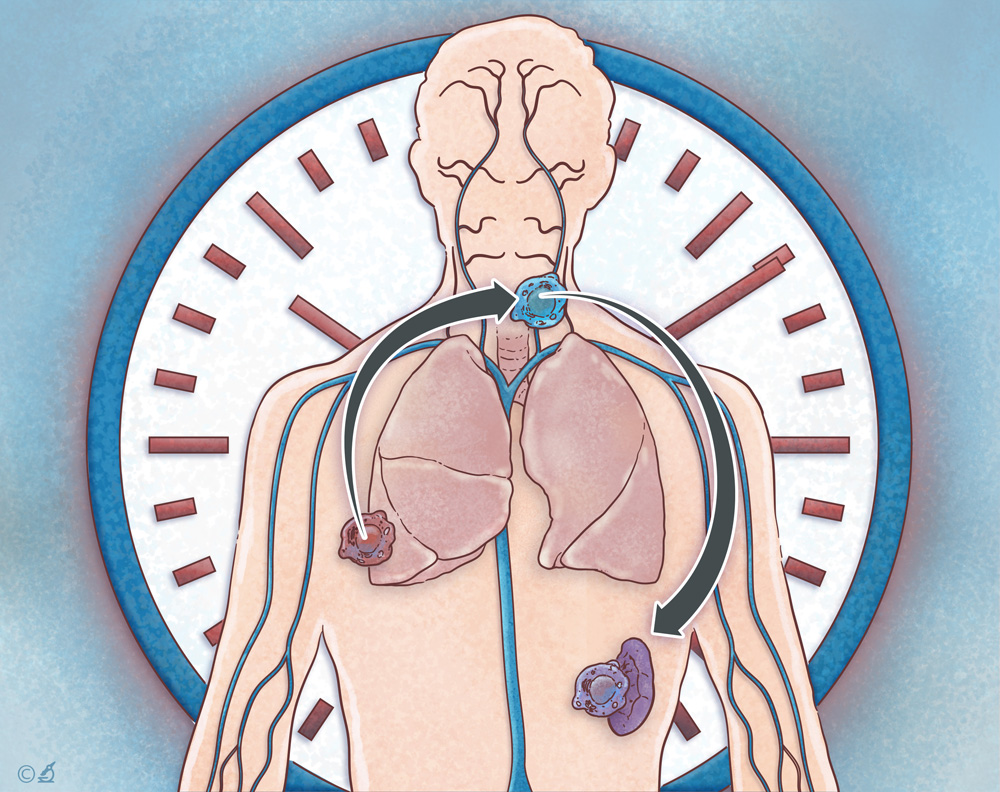
Lineage Tracing of T cell clones
The T cell receptor (TCR) repertoire in each individual is highly diverse and allows for recognition of a wide array of foreign antigens. The TCR also serves as a molecular indicator of cellular origin as well as tumor response, since T cells expressing identical TCRab pairs likely arise from a common cellular ancestor, and T cells responding to tumors proliferate after antigen recognition. Our lab takes advantage of these properties of the TCR to trace the clonal response of individual tumor-specific T cell clones and to separate signals in tumor-specific T cells from noise in ‘bystander’ non-reactive T cells in the same patient. Using single cell technologies, paired with clonal history, we aim to study the origin and evolution of the T cell response during tumor progression, metastasis, and checkpoint blockade:
1) Which T cell types are clonally-expanded in the tumor site, and is clonal expansion linked to the molecular phenotype of each T cell?
2) What is the clonal behavior of tumor-specific T cells over the course of disease and therapy progression?
3) What is the origin of T cell clones that mediate response to checkpoint blockade?