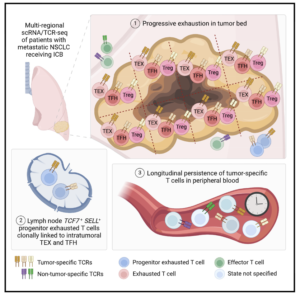
Lineage tracing reveals clonal progenitors and long-
term persistence of tumor-specific T cells during
immune checkpoint blockade
Joy A. Pai, Matthew D. Hellmann, Jennifer L. Sauter, Marissa Mattar, Hira Rizvi, Hyung Jun Woo, Nisargbhai Shah, Evelyn M. Nguyen, Fathema Z. Uddin, Alvaro Quintanal-Villalonga, Joseph M. Chan, Parvathy Manoj, Viola Allaj, Marina K. Baine, Umesh K. Bhanot, Mala Jain, Irina Linkov, Fanli Meng, David Brown, Jamie E. Chaft, Ansuman T. Satpathy
Cancer Cell, 30 March 2023
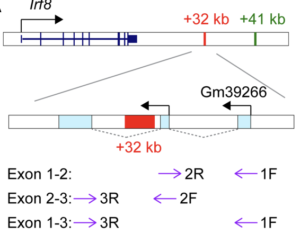
Cis interactions in the Irf8 locus regulate stage-dependent enhancer activation
Tian-Tian Liu, Feiya Ou, Julia A Belk, Prachi Bagadia, David A Anderson 3rd, Vivek Durai, Winnie Yao, Ansuman T Satpathy, Theresa L Murphy, Kenneth M Murphy
Genes & Development, 23 March 2023
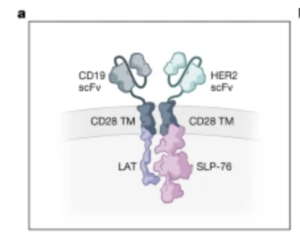
Co-opting signalling molecules enables logic-gated control of CAR T cells
Aidan M Tousley, Maria Caterina Rotiroti, Louai Labanieh, Lea Wenting Rysavy, Won-Ju Kim, Caleb Lareau, Elena Sotillo, Evan W Weber, Skyler P Rietberg, Guillermo Nicolas Dalton, Yajie Yin, Dorota Klysz, Peng Xu, Eva L de la Serna, Alexander R Dunn, Ansuman T Satpathy, Crystal L Mackall, Robbie G Majzner
Nature, 8 March 2023
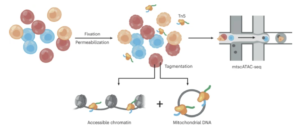
Mitochondrial single-cell ATAC-seq for high-throughput multi-omic detection of mitochondrial genotypes and chromatin accessibility
Caleb A Lareau, Vincent Liu, Christoph Muus, Samantha D Praktiknjo, Lena Nitsch, Pauline Kautz, Katalin Sandor, Yajie Yin, Jacob C Gutierrez, Karin Pelka, Ansuman T Satpathy, Aviv Regev, Vijay G Sankaran, Leif S Ludwig
Nature Protocols, 15 February 2023
A Cre-deleter specific for embryo-derived brain macrophages reveals distinct features of microglia and border macrophages
Simone Brioschi, Julia A Belk, Vincent Peng, Martina Molgora, Patrick Fernandes Rodrigues, Khai M Nguyen, Shoutang Wang, Siling Du, Wei-Le Wang, Gary E Grajales-Reyes, Jennifer M Ponce, Carla M Yuede, Qingyun Li, John M Baer, David G DeNardo, Susan Gilfillan, Marina Cella, Ansuman T Satpathy, Marco Colonna
Immunity, 14 February 2023
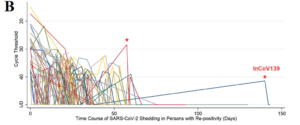
Reinfection with SARS-CoV-2 and Waning Humoral Immunity: A Case Report
Jason D. Goldman, Kai Wang, Katharina Röltgen, Sandra C. A. Nielsen, Jared C. Roach,* ,
Samia N. Naccache, Fan Yang, Oliver F. Wirz, Kathryn E. Yost, Ji-Yeun Lee, Kelly Chun, Terri Wrin,
Christos J. Petropoulos, Inyoul Lee, Shannon Fallen, Paula M. Manner, Julie A. Wallick, Heather A. Algren, Kim M. Murray, Jennifer Hadlock, Daniel Chen, Chengzhen L. Dai, Dan Yuan,
Yapeng Su, Joshua Jeharajah, William R. Berrington, George P. Pappas, Sonam T. Nyatsatsang,
Alexander L. Greninger, Ansuman T. Satpathy, John S. Pauk, Scott D. Boyd, and James R. Heath
Vaccines, 20 December 2022
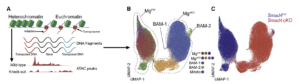
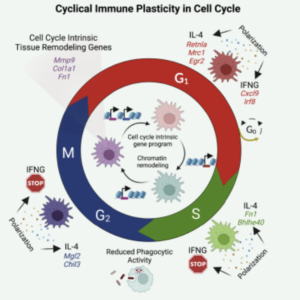
Macrophage inflammatory and regenerative response periodicity is programmed by cell cycle and chromatin state
BenceDaniel, Julia A.Belk, Stefanie L.Meier, Andy Y.Chen, Katalin Sandor, ZsoltCzimmerer, ZsofiaVarga, Krisztian Bene, Frank A.Buquicchio, YanyanQi, Hugo Kitano, Joshua R. Wheeler, Deshka S. Foster, Michael Januszyk, Michael T. Longaker, Howard Y. Chang, Ansuman T. Satpathy
Molecular Cell, 14 December 2022
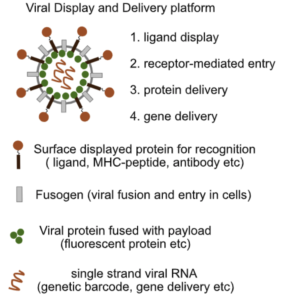
Engineered cell entry links receptor biology with single-cell genomics
Bingfei Yu, Quanming Shi, Julia A. Belk, Kathryn E. Yost, Kevin R. Parker, Rui Li,Betty B. Liu, Huang Huang,
Daniel Lingwood, William J. Greenleaf, Mark M. Davis, Ansuman T. Satpathy, and Howard Y. Chang
Cell, 13 December 2022
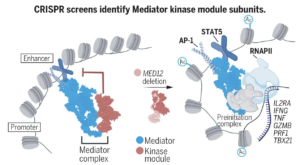
Enhanced T cell effector activity by targeting the Mediator kinase module
Katherine A. Freitas, Julia A. Belk, Elena Sotillo, Patrick J. Quinn, Maria C. Ramello, Meena Malipatlolla, Bence Daniel, Katalin Sandor, Dorota Klysz, Jeremy Bjelajac, Peng Xu, Kylie A. Burdsall, Victor Tieu, Vandon T. Duong,Micah G. Donovan, Evan W. Weber, Howard Y. Chang, Robbie G. Majzner, Joaquin M. Espinosa, Ansuman T. Satpathy, Crystal L. Mackall
Science, 11 November 2022
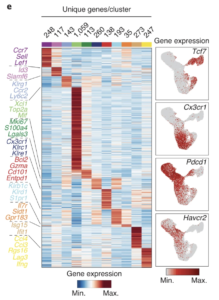
Divergent clonal differentiation trajectories of T cell exhaustion
Bence Daniel, Kathryn E. Yost, Sunnie Hsiung, Katalin Sandor, Yu Xia, Yanyan Qi, Kamir J. Hiam-Galvez, Mollie Black, Colin J. Raposo, Quanming Shi, Stefanie L. Meier, Julia A. Belk, Josephine R. Giles, E. John Wherry, Howard Y. Chang, Takeshi Egawa & Ansuman T. Satpathy
Nature Immunology, 26 October 2022